
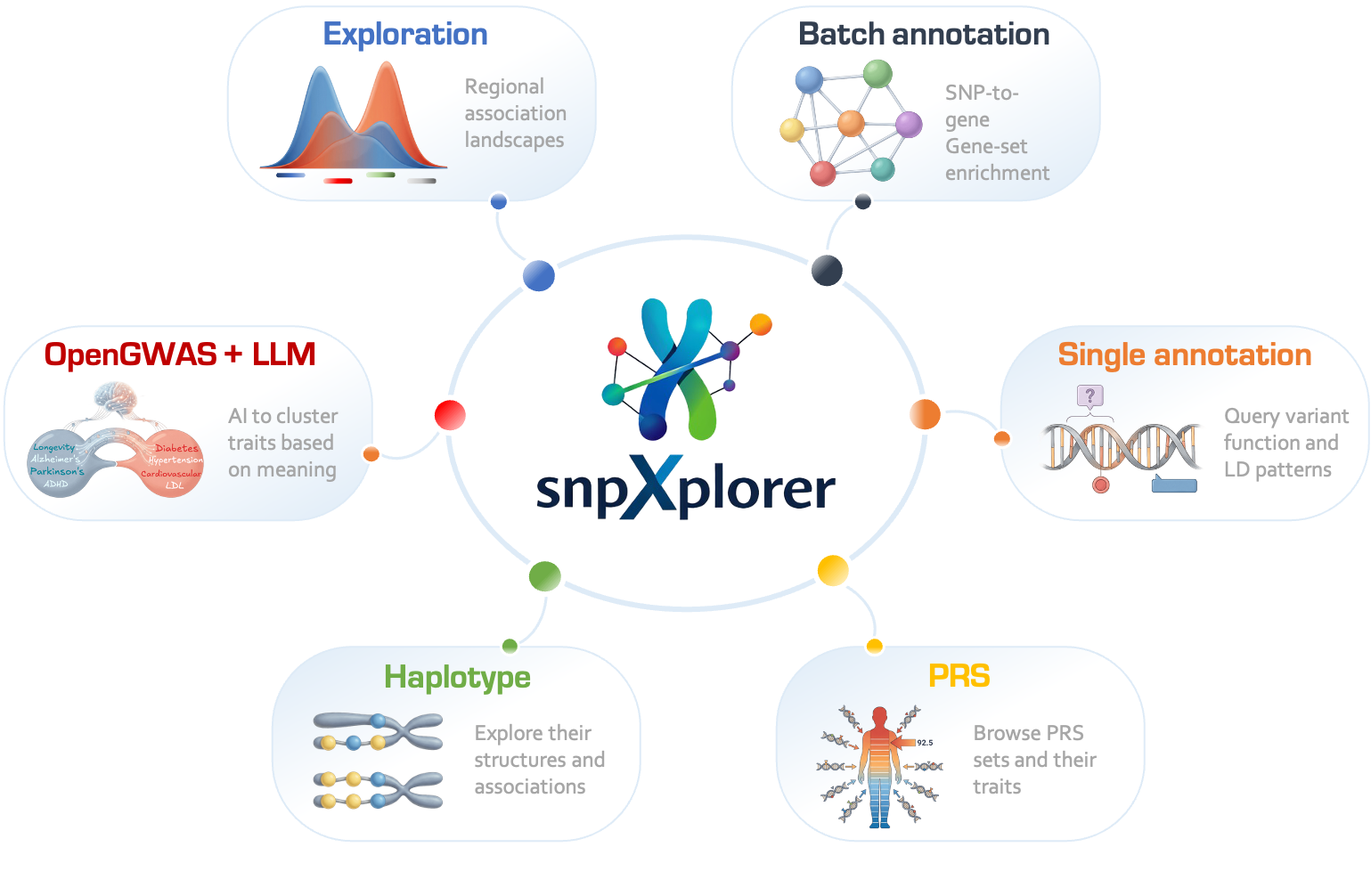
snpXplorer is a free web-server to explore GWAS datasets (Exploration tab) and annotate set of SNPs (Annotation tab).
Exploration
The Exploration tab lets you interactively browse regional association signals across >10,000 GWAS from OpenGWAS. Search a SNP or gene, then overlay multiple studies to compare association landscapes side-by-side.
Customize panels with recombination rates, structural variants (SV), linkage disequilibrium (LD), and haplotypes. Inspect effect sizes, view GTEx expression across >40 tissues, and download the plotted data tables for your own analyses.
Single Annotation
The Single Annotation tab provides a fast, detailed view of one variant at a time. Retrieve predicted consequences (CADD/SIFT/PolyPhen), eQTL/sQTL evidence, LD structure, and GWAS hits from OpenGWAS.
Results are returned for the query variant and its LD partners, with tables you can browse and download. snpXplorer also summarizes the evidence to highlight the most likely affected gene—transparent and data-driven, not magic.
Batch Annotation
The Batch Annotation tab extends Single Annotation to variant lists, enabling high-throughput annotation in one submission. Use it for SNP→gene mapping alone, or include pathway analysis to interpret biology at scale.
For enrichment, genes linked to your variants are tested in a sampling-based framework to identify significantly enriched pathways. Multiple genes can be assigned per variant; submit your list and email, and you’ll receive a complete report.
Haplotypes
The Haplotypes tab supports context-aware exploration beyond single variants. Start from a gene, region, or trait and inspect haplotype definitions, structure, and local association patterns in an interactive workflow.
Because OpenGWAS traits can be highly redundant, snpXplorer groups related traits using semantic similarity derived from large language models. This makes the trait space easier to browse while keeping results transparent and fully downloadable.
PRS
The PRS tab provides variant sets for polygenic risk score construction derived from OpenGWAS studies. We apply clumping and thresholding to obtain approximately independent SNP lists using LD patterns from TopMed.
You can browse a trait, retrieve related traits, and download SNP lists ready for local PRS computation. To protect privacy, genotype data are never uploaded; instead you can run PRS locally with our companion tools jordan or polygenius.
Help & Contacts
The Help, About and contacts tabs describe each module, data sources, and recommended workflows. You can also find references, study information, license information, and the original snpXplorer publication in Nucleic Acid Research.
If snpXplorer supports your research, please cite us. For questions, collaborations, or feedback, email n.tesi@amsterdamumc.nl—we’re happy to help and improve the platform.


